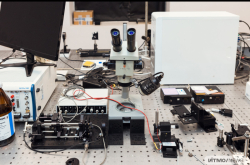
In the recent years, the spread of antibiotic resistance has become a global healthcare issue. As a consequence of excessive antibiotics use in medicine and agriculture, gut microbiota accumulates antibiotic resistance genes (R-genes) in its DNA or metagenome. On the one hand, these genes help regular flora survive. However, on the other hand, recent studies have shown that gut microbiota is capable of sharing resistance genes with pathogens, thus making them resistant to available therapies. In this light, the study of how resistance genes spread becomes especially important.
Programmers from ITMO University and their colleagues from the Research Center of Physical and Chemical Medicine have developed an algorithm called MetaCherchant that makes it possible to explore the environment of a specific gene and see how it changes depending on bacteria species.
“We have created a tool that allows us to identify the difference between the R-genes in the microbial genomes from two or more samples of gut microbiota. We can compare the microbiota of two different people or samples of one person’s microbiota from different time periods – like, for example, before and after antibiotics treatment,” – says Vladimir Ulyantsev, Associate Professor at the Computer Technologies Department at ITMO University – “The resulting data can be visualized and used to theorize as to how a drug-resistant gene is passed on from one species of microbes to another”.
Vladimir Ulyantsev
Studying the environment and the mechanisms of antibiotic R-genes is mainly important for designing effective antimicrobial treatment schemes.
"Using MetaCherchant, we can analyze how microbiota contributes to the spread of resistance to a particular antibiotic class. It is possible that we will be able to predict to which antibiotics the pathogens are more likely to develop a resistance to – and vice-versa. This will help us adjust and fine-tune specific treatments. This is the main task for the next couple of years," says Evgenii Olekhnovich, lead author and researcher at the Center of Physical and Chemical Medicine.
Potential applications of the algorithm are not limited to gut microbiota gene analysis since the program can be also used to study genome samples from soil, water or sewage.
"We can evaluate the spread of resistance within a single bacterial community, such as gut microbiota, as well as between different communities. This allows us, for example, to identify global pathways of how antibiotic resistance spreads through the environment,” says Evgenii Olekhnovich. “The problem of resistance is complex and requires a complex approach, where our tool can be really useful.”
Reference: «MetaCherchant: analyzing genomic context of antibiotic resistance genes in gut microbiota», E. I. Olekhnovich, A. T. Vasilyev, V. I. Ulyantsev et al. Bioinformatics Oct. 28, 2017






